Structure, proteome and genome of Sinorhizobium meliloti phage ΦM5: A virus with LUZ24-like morphology and a highly mosaic genome - ScienceDirect
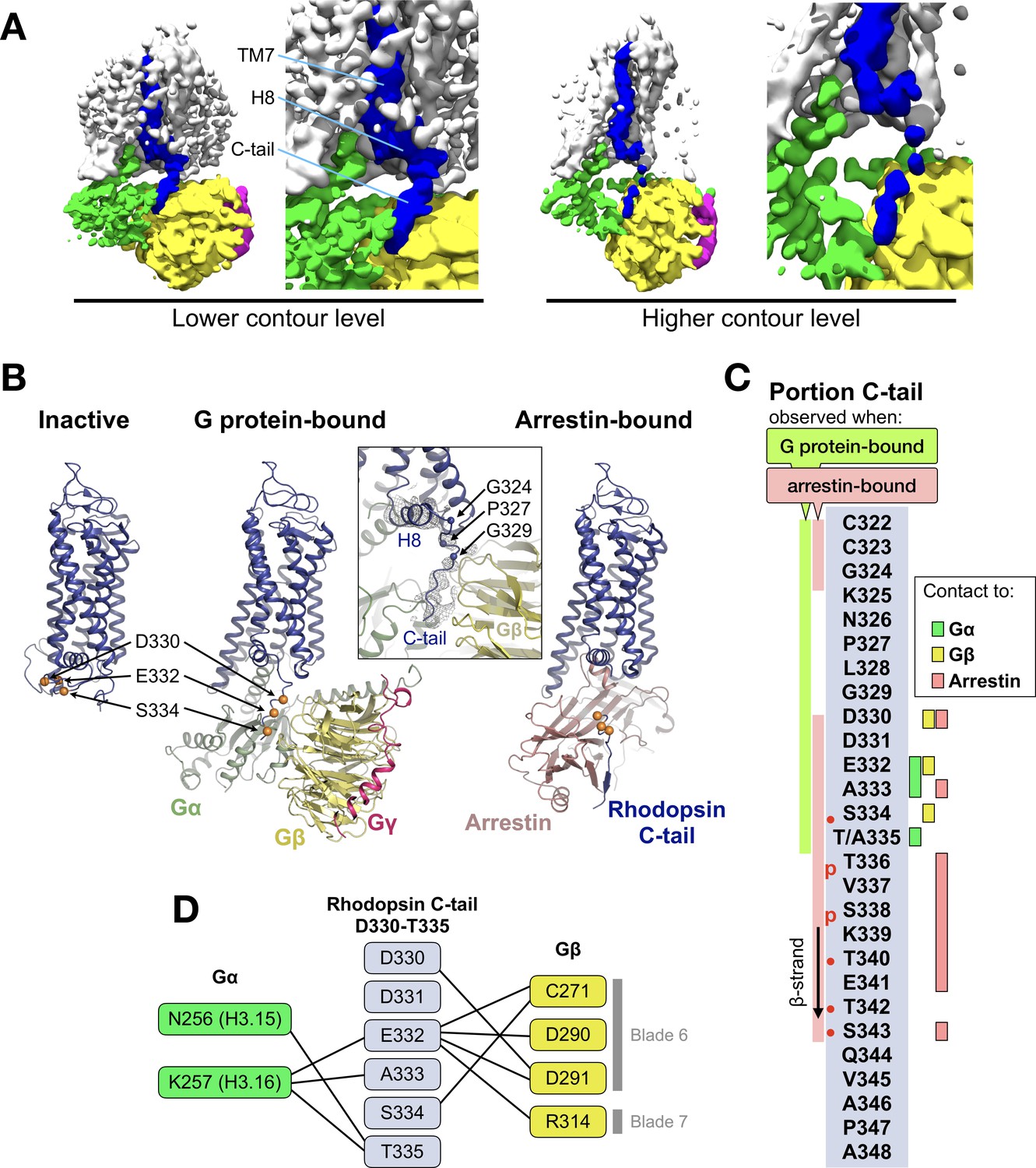
Cryo-EM structure of the rhodopsin-Gαi-βγ complex reveals binding of the rhodopsin C-terminal tail to the gβ subunit | eLife

Cryo–electron microscopy structure and analysis of the P-Rex1–Gβγ signaling scaffold | Science Advances
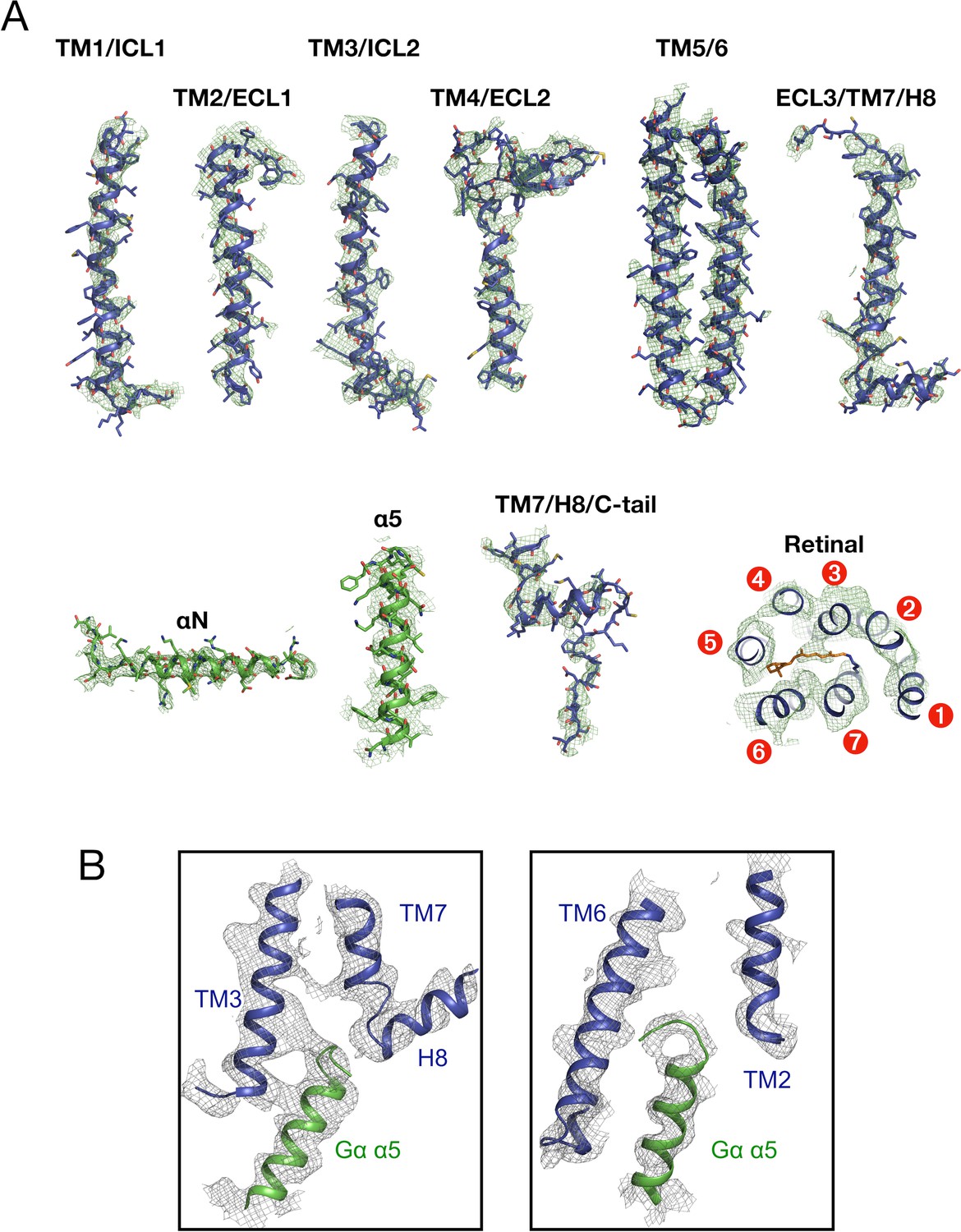
Cryo-EM structure of the rhodopsin-Gαi-βγ complex reveals binding of the rhodopsin C-terminal tail to the gβ subunit | eLife

Crosslinking by ZapD drives the assembly of short, discontinuous FtsZ filaments into ring-like structures in solution | bioRxiv

Crosslinking by ZapD drives the assembly of short, discontinuous FtsZ filaments into ring-like structures in solution | bioRxiv

Crosslinking by ZapD drives the assembly of short, discontinuous FtsZ filaments into ring-like structures in solution | bioRxiv

Cryo–electron microscopy structure and analysis of the P-Rex1–Gβγ signaling scaffold | Science Advances

Crosslinking by ZapD drives the assembly of short, discontinuous FtsZ filaments into ring-like structures in solution | bioRxiv

MemBrain: A Deep Learning-aided Pipeline for Automated Detection of Membrane Proteins in Cryo-electron Tomograms | bioRxiv